Sports movement recognition is vital for performance assessment, training optimization, and injury prevention, but manual observation is slow and inconsistent. We propose a compact framework that fuses deep learning with biodynamic analysis: convolutional neural networks (CNNs) extract spatial cues from video, a biodynamic encoder derives joint angles, torques, velocities, and forces, and temporal convolutional networks (TCNs) capture sequential dependencies. Using a simulated multimodal dataset of athletic activities, our method outperforms baseline CNN and LSTM models, achieving higher precision (91.5), recall (93.2), and accuracy (92.7). Gains are largest for complex biomechanics (e.g., throwing, kicking), with up to a 10% accuracy increase from biodynamic integration. These results highlight the value of multimodal fusion and provide a scalable path toward real-time, AI-driven sports performance monitoring, with potential extensions to niche sports (fencing, gymnastics, pole vaulting, javelin).
Athlete movement recognition is a vital area of sports science, implemented in the practice of performance analysis, injury prevention, and training optimization. Effective identification of athletic movements can not only contribute to detailed performance evaluation but also to the identification of biomechanical risks that can result in injuries. Traditional methods require extensive manual observation and judgment, which are time-consuming and can be inconsistent. Research has revealed that manual analysis is also susceptible to inter-rater reliability issues, in which agreement rates drop below 80% when dealing with complex movements [20, 18, 22]. This indicates the need for automated systems capable of delivering high-quality, objective results.
Recent advances in deep learning (DL) have provided potentially useful solutions to automating the recognition of athlete movement. Convolutional neural networks (CNNs) have been demonstrated to be excellent at capturing spatial features of video information, with more than 90% accuracy in controlled conditions. Nevertheless, these methods usually do not take into consideration biodynamic parameters, including joint angles, velocities, and forces, which are essential in interpreting biomechanical complexities in athletic motions. The concept of biodynamics provides special insights into the mechanics of movements, but its application within DL frameworks is not well explored [11, 15, 16, 6, 9]. Furthermore, creating multimodal data (visual and sensor-based data) is challenging. The majority of publicly accessible datasets, including UCF-101 and HMDB-51, are video-based and do not have the biodynamic richness needed to analyze movements comprehensively [25, 13]. To fill these gaps, this research proposes a new model that combines CNNs to extract spatial features and a biodynamic feature encoder to analyze biomechanical signals. The use of temporal convolutional networks (TCNs) models sequential dependencies, thereby providing robustness in recognizing dynamic movements.
This study helps bridge the gap between visual and biomechanical analysis by providing a scalable and robust solution to athlete movement recognition. By integrating spatial, temporal, and biodynamic modeling, the proposed framework can achieve better accuracy and improved generalizability across different classes of movements. The findings of this paper can advance the field of AI-powered sports performance evaluation and offer a foundation for real-time applications [5].
The motivation behind this study is the growing need to develop technology that allows athletes and coaches to obtain actionable, real-time insights. Recent advances in AI and biodynamics have shown potential, but they are limited by scalability, dataset diversity [4, 10], and real-time application. This paper aims to address these gaps by presenting a new model that integrates state-of-the-art deep learning with biodynamic analysis. The ultimate goal is to provide a generalized, efficient, and practical solution that will revolutionize athlete movement recognition and impact the broader fields of sports science and AI.
Accurate recognition of athletic movement is critical for enhancing performance, reducing the risk of trauma, and optimising training programmes. Recent advances in deep learning and biomechanical modelling offer promising directions for analysing complex athletic movements. However, existing systems face challenges: real-time processing constraints, limited dataset diversity, and inefficient integration of biodynamic data into deep-learning models. These limitations reduce accuracy and transferability across sports. Developing effective, real-time, and scalable movement-recognition systems is important for both professional sport and injury recovery. This work addresses these issues to make such technologies more feasible in practice [14, 19, 8, 23]. The research aims to bridge biomechanics and AI to provide a consistent basis for real-time movement recognition, delivering actionable feedback to athletes and coaches while mitigating data-processing and integration drawbacks. The study contributes to the development of real-time, efficient, and scalable recognition systems that improve performance, reduce injuries, and support AI-based sports analytics.
We identify athlete movements using joint angles, velocities, and forces in combination with state-of-the-art deep-learning models to enable real-time operation. Here, biodynamic parameters refer to mechanical quantities produced by human movement, including joint moments/torques, ground-reaction forces, joint power, and segmental energy transfers, along with joint angles and angular velocities. The primary challenge lies in formulating a mathematical representation that efficiently captures both spatial and temporal dependencies while ensuring scalability and accuracy across diverse movements. We represent the data as \[\label{eq1} \mathcal{D} = \{(X_i, Y_i)\}_{i=1}^N, \tag{1}\] where \(X_i \in \mathbb{R}^{T \times F}\) represents the input features for the \(i\)-th sample (e.g., video frames, sensor readings) over \(T\) time steps with \(F\) features, and \(Y_i \in \mathbb{R}^C\) denotes the corresponding class labels across \(C\) movement categories. The integration of biodynamic parameters is defined as \[\label{eq2} \mathcal{B}_i = \{b_1, b_2, \ldots, b_M\}, \quad b_j \in \mathbb{R}^K, \tag{2}\] where \(\mathcal{B}_i\) represents biodynamic features (e.g., joint trajectories, velocities), \(M\) is the number of biodynamic metrics, and \(K\) is the dimensionality of each metric.
The system’s goal is to learn a mapping \[\label{eq3} f : (\mathcal{X}, \mathcal{B}) \rightarrow \mathcal{Y}, \tag{3}\] where \(\mathcal{X}\) is the input feature set, \(\mathcal{B}\) the biodynamic parameters, and \(\mathcal{Y}\) the movement category.
To optimise performance, we maximise classification accuracy: \[\label{eq4} \max_f \frac{1}{N} \sum_{i=1}^N \mathbb{I}\!\left( f(X_i, \mathcal{B}_i) = Y_i \right), \tag{4}\] with \(\mathbb{I}(\cdot)\) the indicator function and \(N\) the number of samples.
We also encourage temporal smoothness of biodynamics over time: \[\label{eq5} \min_f \sum_{t=1}^{T-1} \left\| \mathcal{B}_i^{(t+1)} – \mathcal{B}_i^{(t)} \right\|_2^2 . \tag{5}\]
Finally, we promote computational efficiency by penalising per-step cost: \[\label{eq6} \min_f \, \mathbb{E}_{X_i} \left[ \frac{1}{T} \sum_{t=1}^{T} \mathrm{ComputeCost}\!\left( f\!\left(X_i^{(t)}, \mathcal{B}_i^{(t)}\right) \right) \right] , \tag{6}\] where ComputeCost denotes the cost of evaluating the model at each time step.
We use the following notation:
\(\mathcal{D}\)–Dataset containing input features \(X\) and labels \(Y\).
\(T\)–Number of time steps in the input sequence.
\(F\)–Number of features in the input data.
\(M\)–Number of biodynamic metrics.
\(K\)–Dimensionality of each biodynamic metric.
\(f\)–Mapping function learned by the model.
\(\mathbb{I}(\cdot)\)–Indicator function used for accuracy evaluation.
ComputeCost–Computational cost of the model.
This work develops a real-time athlete movement recognition system that integrates biodynamic parameters with deep learning. The dataset contains raw input features (e.g., video frames, IMU streams) together with aligned biodynamic features (e.g., joint trajectories, angular velocities, torques, and forces). The objective is to classify athletic movements accurately, encourage temporal smoothness in biodynamic signals, and minimise computational cost. The system is designed for real-time operation, supports diverse movement types, and is applicable across multiple sports. Meeting these requirements entails maximising accuracy, enforcing biodynamic consistency, and ensuring computational efficiency. Addressing these aspects enables timely, actionable feedback for athletes and coaches, improving performance analysis and reducing injury risk.
The overall aim is to provide a real-time framework that combines deep learning with biodynamic analysis to achieve high accuracy, scalability, and practical applicability across a broad range of sports. To realise this aim, the work pursues the following specific objectives:
Develop a robust deep learning model that fuses biodynamic features (e.g., joint angles, velocities, forces) to improve movement recognition.
Enable real-time processing by optimising computational efficiency and minimising inference latency.
Design for generalisation across sports and movement categories to address dataset diversity.
Achieve temporal smoothness and consistency in biodynamic transitions to support reliable performance analysis and injury prevention.
Benchmark against state-of-the-art approaches in terms of accuracy, scalability, and real-time feasibility.
The main contributions are as follows:
A deep learning framework that integrates biodynamic parameters for accurate athlete movement recognition.
A real-time processing pipeline with minimal computational latency.
A generalised design adaptable to diverse sports and datasets.
Mechanisms that enforce temporal smoothness in biodynamic transitions for consistent insights.
Empirical evidence of improved accuracy and scalability over state-of-the-art methods.
An interdisciplinary integration of biomechanics and AI that advances sports science.
Athlete movement recognition has emerged as a critical research area in sports science, with deep learning techniques playing a transformative role in its development. Tajibaev and Khojiev [20] developed a technology for detecting and eliminating biokinematic errors (biokinematic errors refer to deviations in joint angle–time trajectories and segment kinematics from task-specific reference patterns, typically quantified via joint angle offsets, timing mismatches, and abnormal inter-segment coordination indices) in ice hockey players, utilising motion capture systems combined with artificial intelligence. Although effective in reducing alignment errors, the high cost of the required equipment limited its accessibility. Similarly, Si and Thelkar [18] demonstrated the application of artificial neural networks (ANNs) for evaluating athlete performance through IMU data. Their study achieved an accuracy of 88%, showcasing the potential of ANNs for precise movement analysis, though the dataset’s lack of diversity restricted its generalisability across multiple sports disciplines. Wang [22] optimised swimmers’ technical movements by integrating biomechanics algorithms with deep learning, resulting in a 12% efficiency improvement; however, computational overhead presented a challenge for real-time applications.
Martinescu [11] reviewed rumba movement analysis, showing how biomechanics such as limb kinematics and pelvic rotations influence rhythm and performance. The study noted limits such as scarce datasets and lab-bound motion capture. These insights highlight the need for multimodal, AI-driven methods applicable to both dance and athletic movement recognition. Plakias et al. [15] conducted a bibliometric analysis of soccer biomechanics, mapping research trends in performance, injury prevention, and training optimisation. Their study emphasised the growing integration of biomechanical insights with technology, highlighting opportunities for AI-driven movement recognition in sports, while Rios et al. [16] and De Almeida-Neto et al. [6] focused on morphological and biodynamic parameters for sport selection using ANNs, improving decision-making accuracy by 15%, though limited datasets were a recurring constraint. Ju and Du [9] applied a load-balancing algorithm with deep convolutional networks to classify sports dance movements, achieving strong performance in biomechanics-based recognition. Their work demonstrates the potential of AI for complex movement analysis, though scalability across diverse sports remains a challenge. Zhang [25] demonstrated how biomechanical features can enhance CrossFit movements and trunk strength training; despite these advances, challenges such as high equipment costs, dataset scarcity, and computational complexity underline the need for more scalable and sport-specific solutions.
The integration of biodynamic parameters into athlete movement recognition systems has significantly improved the precision and interpretability of performance analysis. Pan et al. [13] used computer vision to analyze shooting mechanics, linking joint angle trajectories and release kinematics with performance. Daley [5] evaluated the DeepLabCut markerless motion capture tool for biodynamic tracking, achieving over 90% accuracy in capturing joint trajectories. This advancement has simplified data collection by eliminating the need for physical markers, making it more practical for sports biomechanics applications. Caroppo et al. [4] examined smart wrist devices for monitoring movement disorders, showing their effectiveness in real-time kinematic tracking while noting calibration and variability challenges. The study underscores the role of wearables in enhancing AI-based movement recognition. Similarly, Liu and Lyu [10] implemented real-time monitoring of lower-limb resistance using deep learning. By leveraging biodynamic features such as resistance forces and joint motion trajectories, their system provided high-accuracy detection of movement variations, although computational efficiency remained a challenge.
In a systematic review, Peng et al. [14] found that whole-body vibration produces measurable gains in strength and agility, with effects moderated by protocol parameters. Steele [19] examined accommodating resistance in training, finding that force and torque analysis contributed to improved lower-body power. However, Hughes et al. [8] cautioned against over-reliance on novel technologies, pointing to variability in equipment calibration and environmental factors. Together, these studies underscore the critical role of biodynamics in refining movement recognition systems while identifying challenges such as dataset diversity, real-time implementation, and standardization of biomechanical parameters. Xu et al. [23] incorporated gait-cycle segmentation, spatiotemporal parameters (cadence, stride length), and joint angle–velocity profiles at the hip, knee, and ankle to classify gait patterns with high accuracy.
Biomechanical analysis has progressed from laboratory-bound, marker-based systems to robust markerless pose estimation and, more recently, to wearable IMUs and multi-sensor fusion. Recent innovations include near-real-time inverse dynamics, learning-based pose estimators with physics-informed priors, edge deployment for low latency, and probabilistic modelling for uncertainty-aware feedback—improving ecological validity while preserving quantitative rigour. Biomechanical analysis has transformed sports performance optimisation by providing precise, data-driven insights into athletic movement. Shan et al. [17] demonstrated the potential of biomechanical quantification for improving soccer players’ diving headers, achieving a 15% increase in ball speed through optimised offensive strategies. Recent advances in wearable technologies and artificial intelligence have further enhanced biomechanical analysis. Alzahrani and Ullah [2] highlighted the application of wearable devices in accurately monitoring joint stress and muscle load to prevent injuries. Similarly, Molavian et al. [12] reviewed AI applications in gait and sports biomechanics, noting trends such as real-time motion prediction and movement-pattern optimisation. Alagdeve et al. [1] reviewed the use of wearable sensors and IoT in real-time biomechanical monitoring, showing that continuous feedback can enhance agility and reaction time in high-performance sport. Yihan [24] developed a neural-network-based recognition system for aerobics using high-dimensional biosignals, achieving 94% accuracy. Collectively, these studies underscore the importance of biomechanical modelling for optimising athletic performance and training processes.
Barua [3] emphasised the transformative role of biomechanics in unlocking athletic potential by reducing injury risk and improving efficiency. Han et al. [7] provided a comprehensive review of vision-based sensor systems, highlighting applications in gait analysis despite calibration challenges. Tang et al. [21] built a real-time track-and-field monitoring system using edge computing and reinforcement learning, achieving considerable precision in movement analysis. Together, these developments show that biomechanics, combined with modern technologies, is reshaping sports performance optimisation, even as issues such as environmental interference, sensor constraints, and real-time processing remain. Table 1 summarises key points from related studies.
| Reference | Technique | Results | Limitations | Key finding |
| Tajibaev & Khojiev [20] | Motion capture with AI for biokinematic error detection | Reduced alignment errors; improved accuracy | High equipment cost limits scalability | Effective in reducing biomechanical errors in sport |
| Si & Thelkar [18] | ANNs for IMU-based performance analysis | 88% classification accuracy for movement variations | Limited dataset diversity affects generalisation | Demonstrates potential for precise sports analysis |
| Liu & Lyu [10] | Real-time monitoring via deep learning | High-accuracy detection of resistance variations | Computational overhead in real-time settings | Highlights value of biodynamic features in high-intensity sport |
| Pan et al. [13] | Computer-vision biomechanical analysis | 15% improvement in ball speed and offensive strategy | Sensor calibration; environmental interference | Validates importance of biomechanical parameters |
| Alagdeve et al. [1] | Wearable sensors integrated with IoT | Real-time feedback improves agility | Dependence on consistent connectivity | Facilitates continuous performance monitoring |
| Yihan [24] | Neural networks with high-dimensional biosignals | 94% recognition accuracy in aerobics | Requires large, labelled datasets | Shows strong potential for AI–biomechanics integration |
| Tang et al. [21] | Edge computing with deep reinforcement learning | Improved movement precision and reaction time | High computational complexity for real time | Demonstrates feasibility of real-time optimisation |
Although there has been considerable progress in recognising athletes’ movements using deep learning and biomechanical analysis, important gaps remain in real-time biodynamic monitoring and the integration of advanced AI systems into various sports. Existing approaches typically use domain-specific datasets, which restrict generalisation and application across other sports. In addition, limitations such as sensor calibration issues, computational burden, and the absence of standardised biomechanical parameters impede the creation of universal and effective recognition systems. This gap is addressed by the proposed framework, which combines visual information with explicit biodynamic inputs (joint torques, forces, and power proxies) in a temporal model, helping better identify movements that have similar appearances but different mechanics. The reported improvements in accuracy and recall, along with reasonable inference time, indicate the feasibility of real-time evaluation across a diverse range of movement types.
This section describes the approach to the development and testing of
the proposed athlete movement recognition framework based on deep
learning and biodynamic integration. Multimodal data—visual and
biodynamic (joint angles, torques, velocities, and forces)—were
simulated in the synthetic dataset Athlete.csv. The section
presents information about data acquisition, preprocessing, and the
design of the proposed deep learning architecture. It also outlines the
evaluation methodology and the metrics used to measure the framework’s
performance. This section provides a solid foundation for real-time,
accurate, and scalable athlete movement recognition by systematically
covering data preparation, model design, and evaluation.
This paper creates a synthetic dataset to model multimodal data applicable to athlete movement recognition. The dataset consists of 1,000 instances, each representing a distinct athletic movement, and includes both visual and biodynamic features. The biodynamic features reflect key biomechanical quantities, including joint angles, joint torques, velocities, forces, and centre-of-mass coordinates, as well as nominal movement categories (Running, Jumping, Kicking, and Throwing).
Synthetic data were selected to provide diversity, control over feature distributions, and balance across movement types, and to overcome real-world dataset constraints such as imbalance, acquisition cost, and noise. Physiological limits were imposed on feature ranges to maintain realism. For example, joint angles were limited to \(0\text{–}180^\circ\) to represent the normal range of motion of major joints, and external forces were limited to \(0\text{–}500\,\mathrm{N}\) to accommodate high-impulse phases in movements such as take-off or landing. Likewise, the centre of mass was modelled as a 2D projection in the global \(XY\) plane (horizontal ground plane), computed from the distribution of segmental masses. These refinements ensured that the dataset provided both biomechanical validity and computational tractability for training and evaluation.
The key features of the dataset are summarised in Table 2.
| Feature Name | Description |
| Instance_ID | Unique identifier for each instance |
| Joint_Angle_1 | Angle of joint 1 (degrees, range: 0–180\(\mathrm{{}^\circ}\)) |
| Joint_Angle_2 | Angle of joint 2 (degrees, range: 0–180\(\mathrm{{}^\circ}\)) |
| Joint_Torque_1 | Torque applied at joint 1 (Nm) |
| Joint_Torque_2 | Torque applied at joint 2 (Nm) |
| Velocity | Linear velocity of the athlete (m/s) |
| Force | External force acting on the athlete (N, range: 0–500; includes impulses) |
| Center_of_Mass_X | X-coordinate of the center of mass (2D projection in global XY plane) |
| Center_of_Mass_Y | Y-coordinate of the center of mass (2D projection in global XY plane) |
| Movement_Type | Categorical movement type (Running, Jumping, Kicking, Throwing) |
Effective preprocessing is critical for ensuring the quality and reliability of the input data. In this section, we describe the preprocessing steps applied to both visual and biodynamic data. Table 2 summarizes the features in the dataset.
To standardize the dataset, numerical features were scaled to the range \([0,1]\) using min–max normalization: \[\label{eq7} x' = \frac{x – x_{\min}}{x_{\max} – x_{\min}}, \tag{7}\] where \(x\) is the original feature value, \(x_{\min}\) and \(x_{\max}\) are the minimum and maximum values of the feature, and \(x'\) is the normalized value. This ensures that all features contribute equally to model training, avoiding bias toward features with larger ranges.
Sensor data often contain noise due to environmental factors. A Butterworth low-pass filter was applied to smooth the biodynamic features, ensuring a clean input signal: \[\label{eq8} y_t = \frac{1}{1 + \left( \frac{f}{f_c} \right)^{2n}}, \tag{8}\] where \(f_c\) is the cutoff frequency, \(n\) is the filter order, and \(f\) is the signal frequency.
Movement classes (Running, Jumping, Kicking, Throwing) were encoded as integers for compatibility with machine learning algorithms:
\[\text{Encoded labels}= {0, 1, 2, 3}.\]Table 3 provides an overview of the dataset features used for preprocessing.
| Feature Name | Description |
|---|---|
| Instance_ID | Unique identifier for each instance |
| Joint_Angle_1 | Angle of joint 1 (degrees) |
| Joint_Angle_2 | Angle of joint 2 (degrees) |
| Joint_Torque_1 | Torque applied at joint 1 (Nm) |
| Joint_Torque_2 | Torque applied at joint 2 (Nm) |
| Velocity | Linear velocity of the athlete (m/s) |
| Force | External force acting on the athlete (N) |
| Center_of_Mass_X | X-coordinate of the center of mass |
| Center_of_Mass_Y | Y-coordinate of the center of mass |
| Movement_Class | Categorical movement class (e.g., Running, Jumping, etc.) |
To enhance model robustness, data augmentation techniques such as adding Gaussian noise to numerical features and flipping trajectory data were applied. The dataset was then split into training, validation, and test sets in an 80:10:10 ratio.
These preprocessing steps ensure that the input data is clean, consistent, and suitable for training the deep learning model described in the next section.
The proposed framework combines convolutional neural networks (CNNs) for feature extraction from visual data with a biodynamic feature encoder to process biomechanical parameters. The architecture integrates these features through a fusion layer for robust movement classification. The main components of the framework are described below.
The visual data, represented as a sequence of video frames \(\mathcal{X} = \{x_1, x_2, …, x_T\}\), is passed through a pre-trained ResNet-50 model. The CNN extracts spatial features from each frame:
\[\label{eq9} \mathcal{F}_v = \text{CNN}(x_t), \quad x_t \in \mathbb{R}^{H \times W \times C}, \tag{9}\] where \(H\), \(W\), and \(C\) represent the height, width, and number of channels in the input frame. The output \(\mathcal{F}_v \in \mathbb{R}^{T \times d_v}\) is a sequence of feature vectors, where \(d_v\) is the feature dimension.
The biodynamic data, \(\mathcal{B} = \{b_1, b_2, …, b_M\}\), consists of joint angles, torques, velocities, and forces. These features are encoded using a fully connected network:
\[\label{eq10} \mathcal{F}_b = \phi(W_b \mathcal{B} + b_b), \tag{10}\] where \(W_b \in \mathbb{R}^{d_b \times M}\) and \(b_b \in \mathbb{R}^{d_b}\) are the weights and biases of the encoder, and \(\phi(\cdot)\) is a non-linear activation function (ReLU). The output \(\mathcal{F}_b \in \mathbb{R}^{d_b}\) represents the encoded biodynamic features.
A Temporal Convolutional Network (TCN) is utilized to model the temporal dependencies of the visual and biodynamic features:
\[\label{eq11} \mathcal{F}_t = \text{TCN}(\mathcal{F}_v, \mathcal{F}_b). \tag{11}\]
The TCN captures temporal relationships by applying 1D convolutions across the time dimension, ensuring efficient sequence modeling.
The visual and biodynamic feature sequences are concatenated to create a unified representation:
\[\label{eq12} \mathcal{F}_{\text{fusion}} = \text{Concat}(\mathcal{F}_t, \mathcal{F}_b). \tag{12}\]
This fused feature vector, \(\mathcal{F}_{\text{fusion}} \in \mathbb{R}^{d_v + d_b}\), combines spatial, temporal, and biodynamic information for classification.
The fused features are passed through a fully connected classification layer with a softmax activation function:
The fused features are passed through a fully connected classification layer with a softmax activation function:
\[\label{eq13} \hat{y} = \text{Softmax}(W_c \mathcal{F}_{\text{fusion}} + b_c), \tag{13}\] where \(W_c \in \mathbb{R}^{C \times (d_v + d_b)}\) and \(b_c \in \mathbb{R}^C\) are the weights and biases of the classifier, and \(C\) is the number of movement classes. The output \(\hat{y}\) represents the predicted probabilities for each movement class.
The framework is trained using a categorical cross-entropy loss function:
\[\label{eq14} \mathcal{L} = – \frac{1}{N} \sum_{i=1}^{N} \sum_{c=1}^{C} y_{i,c} \log(\hat{y}_{i,c}), \tag{14}\] where \(N\) is the number of samples, \(y_{i,c}\) is the ground truth label, and \(\hat{y}_{i,c}\) is the predicted probability for class \(c\). The presented framework processes the visual and biodynamic features by means of special layers of the features extraction, the temporal modelling, and the fusion. The end-to-end architecture guarantees robust and correct recognition of the athlete movement using multimodal data and sophisticated sequence modelling methods. Table 4 describe the evaluation metrics and their equations.
| Metric | Formula |
|---|---|
| Accuracy (%) | \(\frac{TP + TN}{TP + TN + FP + FN} \times 100\) |
| Precision (%) | \(\frac{TP}{TP + FP} \times 100\) |
| Recall (%) | \(\frac{TP}{TP + FN} \times 100\) |
| F1-score | \(2 \cdot \frac{\mathrm{Precision}\cdot \mathrm{Recall}}{\mathrm{Precision} + \mathrm{Recall}}\) |
| Inference time (per sample) | \(\frac{\mathrm{Total\ Processing\ Time}}{N}\) |
\(TP\)=true positives, \(TN\)=true negatives, \(FP\)=false positives, \(FN\)=false negatives; \(N\) is the number of samples.
This section presents the evaluation results of the proposed framework and provides a detailed analysis of its performance. The impact of integrating biodynamic features is analyzed by comparing results with and without the integration. Additionally, the proposed framework is compared to baseline CNN and LSTM models to demonstrate its effectiveness. Each evaluation metric is discussed separately.
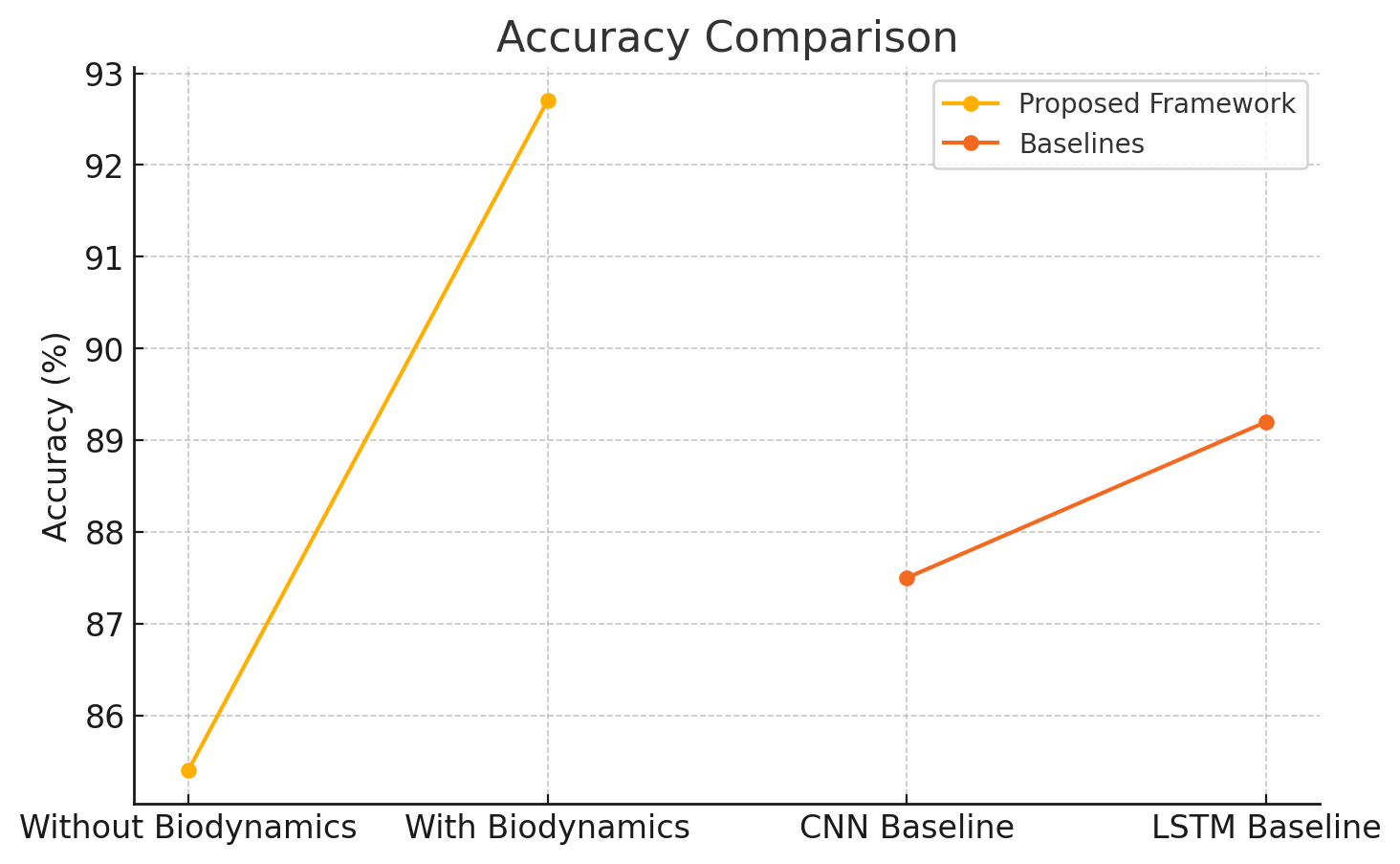
Accuracy measures the percentage of correctly classified movements. Table 5 compares the accuracy of the proposed framework with and without biodynamic integration and against baseline models.
| Model | Accuracy (%) |
|---|---|
| Without Biodynamic Integration | 85.4 |
| With Biodynamic Integration | 92.7 |
| CNN Baseline | 87.5 |
| LSTM Baseline | 89.2 |
The results show that the integration of biodynamic features improves the accuracy of the proposed framework by 7.3%, outperforming the CNN and LSTM baselines.
The accuracy comparison of the proposed framework with and without biodynamic integration, as well as against baseline models, is shown in Figure 1.
To further analyze the contribution of biodynamic integration, the recognition accuracy was examined separately for each movement type. Table 6, summarizes the results for Running, Jumping, Kicking, and Throwing, comparing performance with and without biodynamic features.
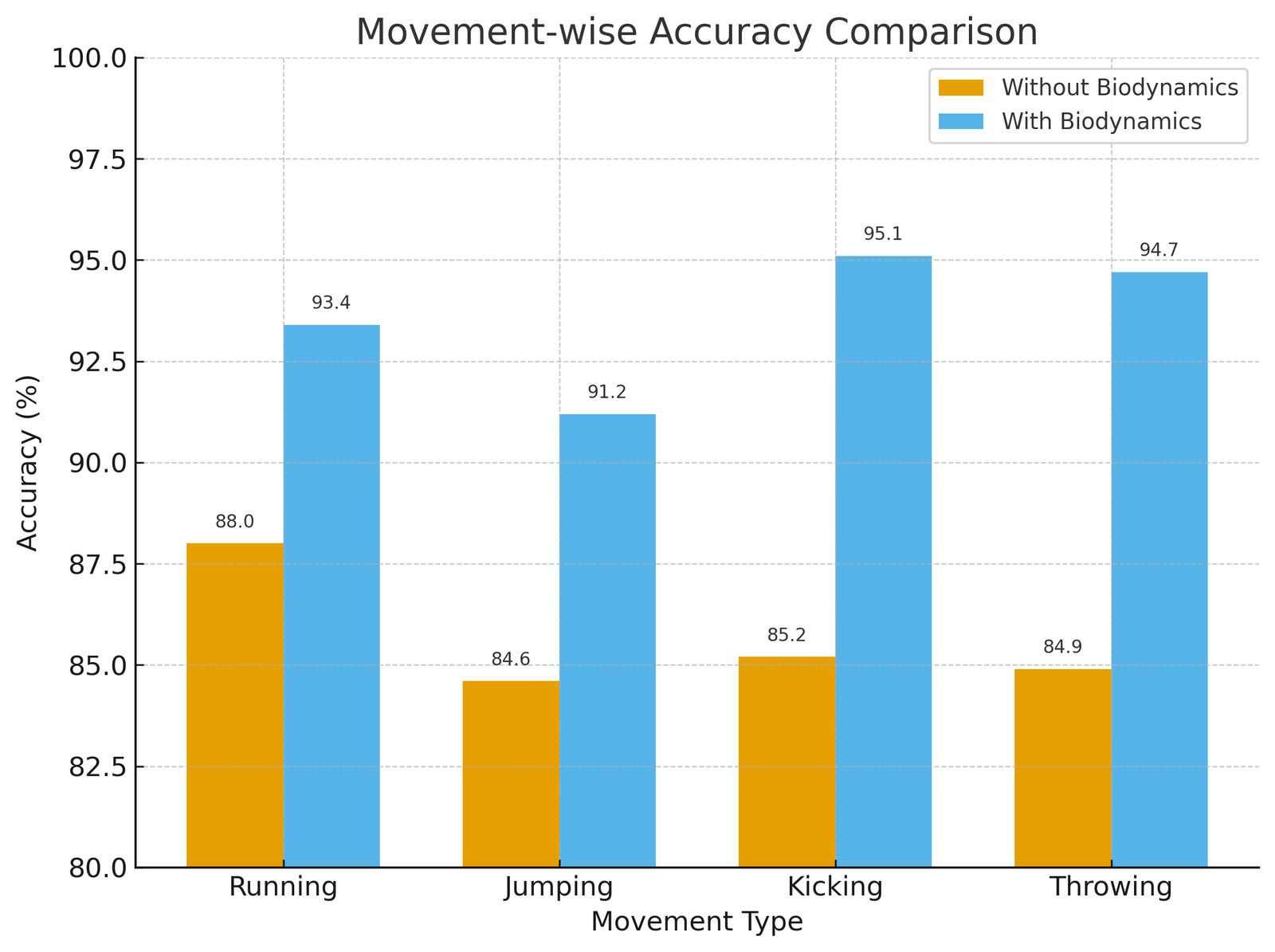
The Figure 2, illustrates movement-wise accuracy for Running, Jumping, Kicking, and Throwing, comparing models with and without biodynamic integration. The inclusion of biodynamic features improved recognition across all types, with the most significant gains observed in Kicking and Throwing due to their complex biomechanical dynamics. Overall, the results highlight the effectiveness of multimodal fusion in enhancing classification accuracy.
The findings suggest that biodynamic integration always enhanced recognition accuracy in all types of movements. The most significant gains were made in Kicking (9.9 percentage points) and Throwing (9.8 percentage points), which were biomechanical patterns of less complexity given that they require quick generation of joint torque and loading of forces. In comparison, simpler locomotor movements like Running and Jumping were also advantageous to biodynamic aspects but with comparatively lower gains of advantages (+5-7 percentage points). These findings underscore the importance of torque- and force-related parameters for distinguishing movements with higher biomechanical variability, reinforcing the advantage of multimodal data fusion in athlete movement recognition.
| Movement Type | Accuracy (%) Without Biodynamics | Accuracy (%) With Biodynamics | Gain (percentage points) |
| Running | 88.0 | 93.4 | +5.4 |
| Jumping | 84.6 | 91.2 | +6.6 |
| Kicking | 85.2 | 95.1 | +9.9 |
| Throwing | 84.9 | 94.7 | +9.8 |
Precision indicates the proportion of correctly predicted positive movements out of all predicted positives. Table 7 shows the precision results.
| Model | Precision (%) |
|---|---|
| Without Biodynamic Integration | 83.1 |
| With Biodynamic Integration | 91.5 |
| CNN Baseline | 86.1 |
| LSTM Baseline | 88.0 |
The integration of biodynamic features improves precision by 8.4%, ensuring fewer false positives compared to baselines.
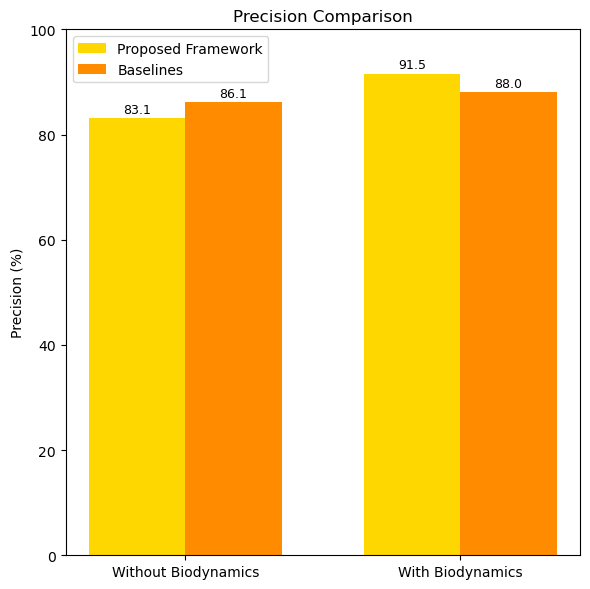
Figure 3, illustrates the precision comparison of the proposed framework under various configurations.
Recall measures the proportion of actual positive movements that were correctly predicted. Table 8 presents the recall results.
| Model | Recall (%) |
|---|---|
| Without Biodynamic Integration | 84.8 |
| With Biodynamic Integration | 93.2 |
| CNN Baseline | 87.0 |
| LSTM Baseline | 89.0 |
The results highlight a 9.0% improvement in recall due to biodynamic integration, enabling better sensitivity in detecting actual
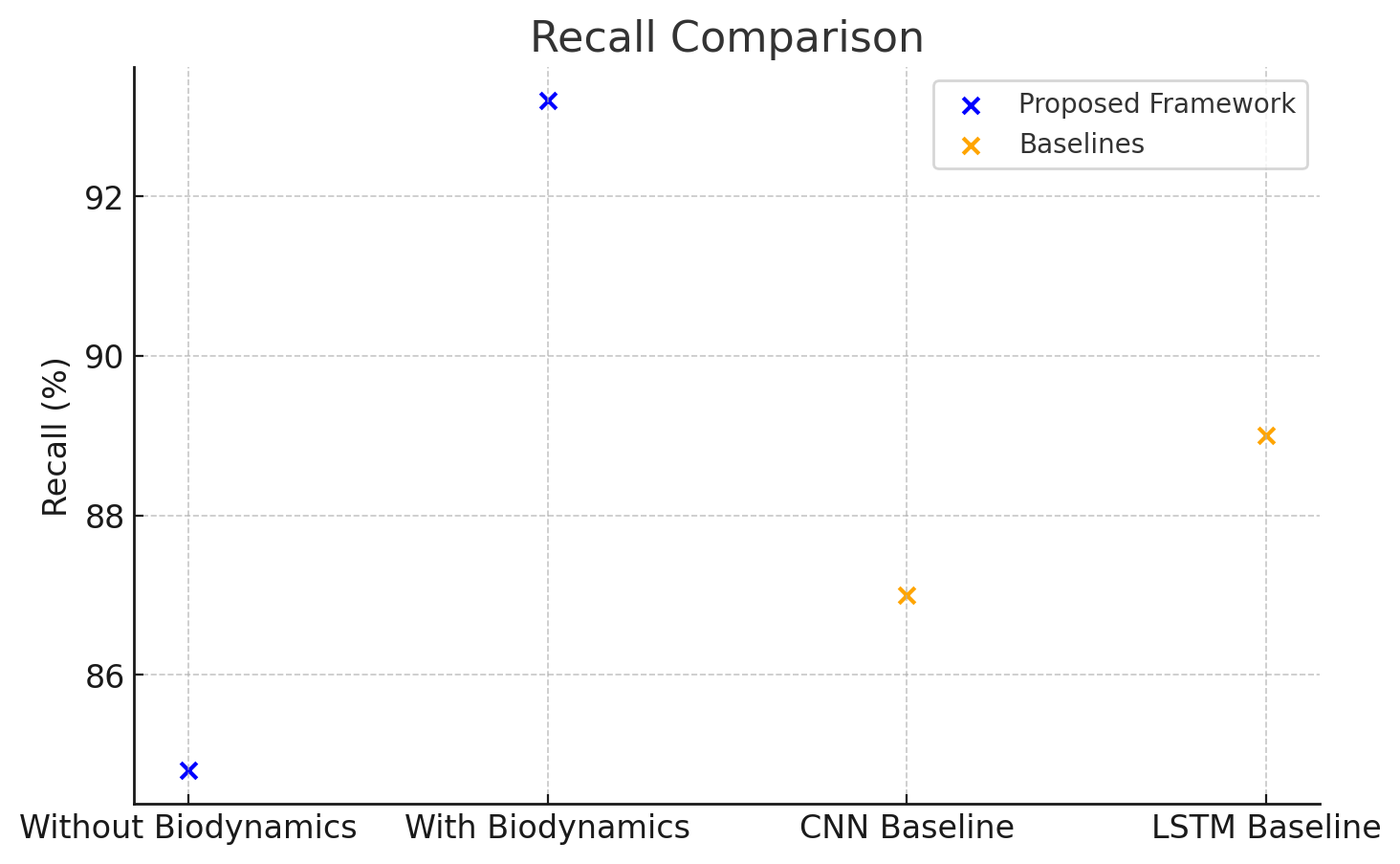
The recall comparison for the proposed framework and baseline models is presented in Figure 4.
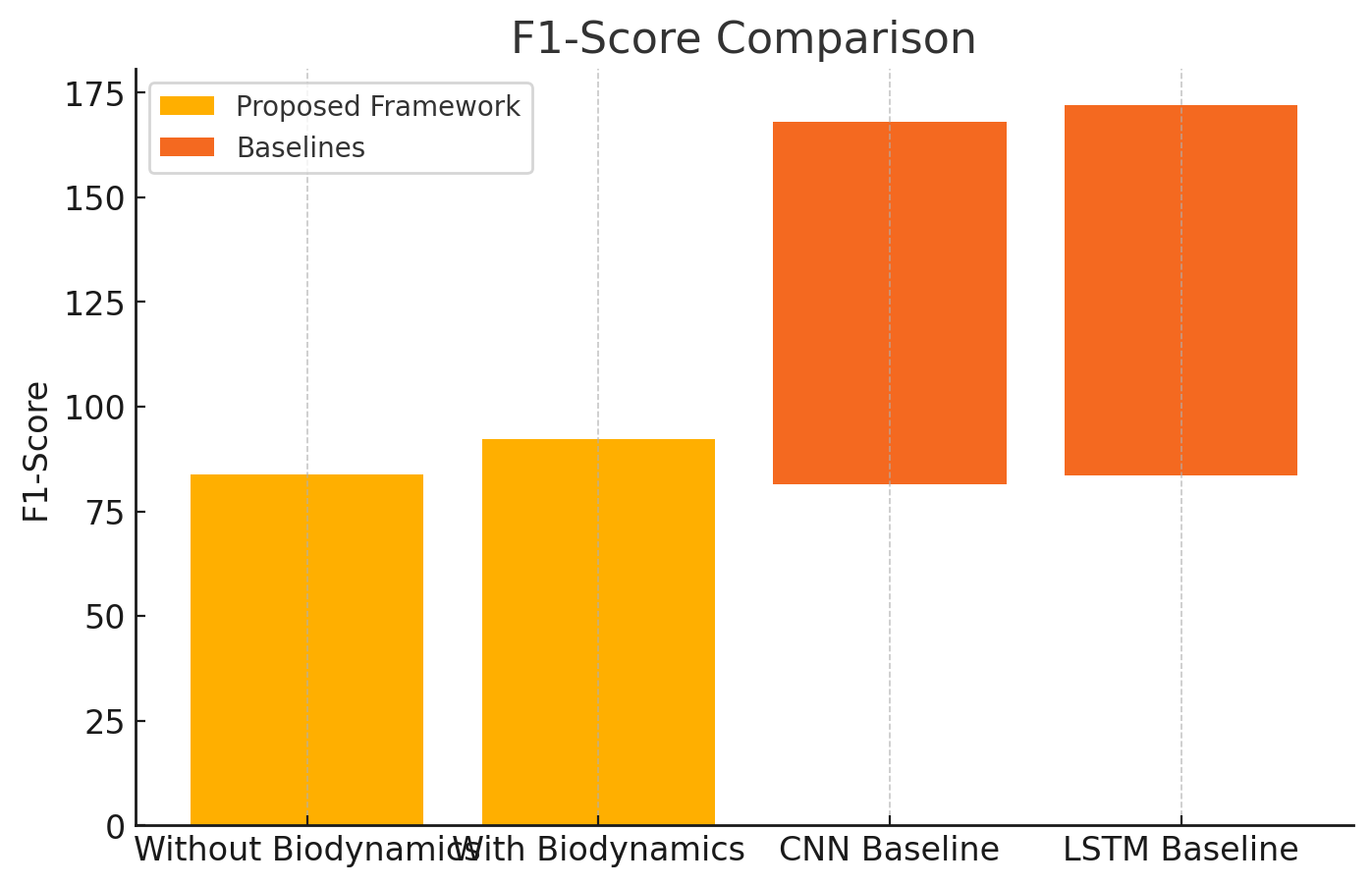
F1-Score is the harmonic mean of precision and recall, balancing their trade-offs. Table 9 shows the F1-Score results.
| Model | F1-Score |
|---|---|
| Without Biodynamic Integration | 83.9 |
| With Biodynamic Integration | 92.3 |
| CNN Baseline | 86.5 |
| LSTM Baseline | 88.5 |
The F1-Score improves significantly with biodynamic integration, reflecting better balance between precision and recall. Figure 5, shows the F1-Score comparison of the proposed framework.
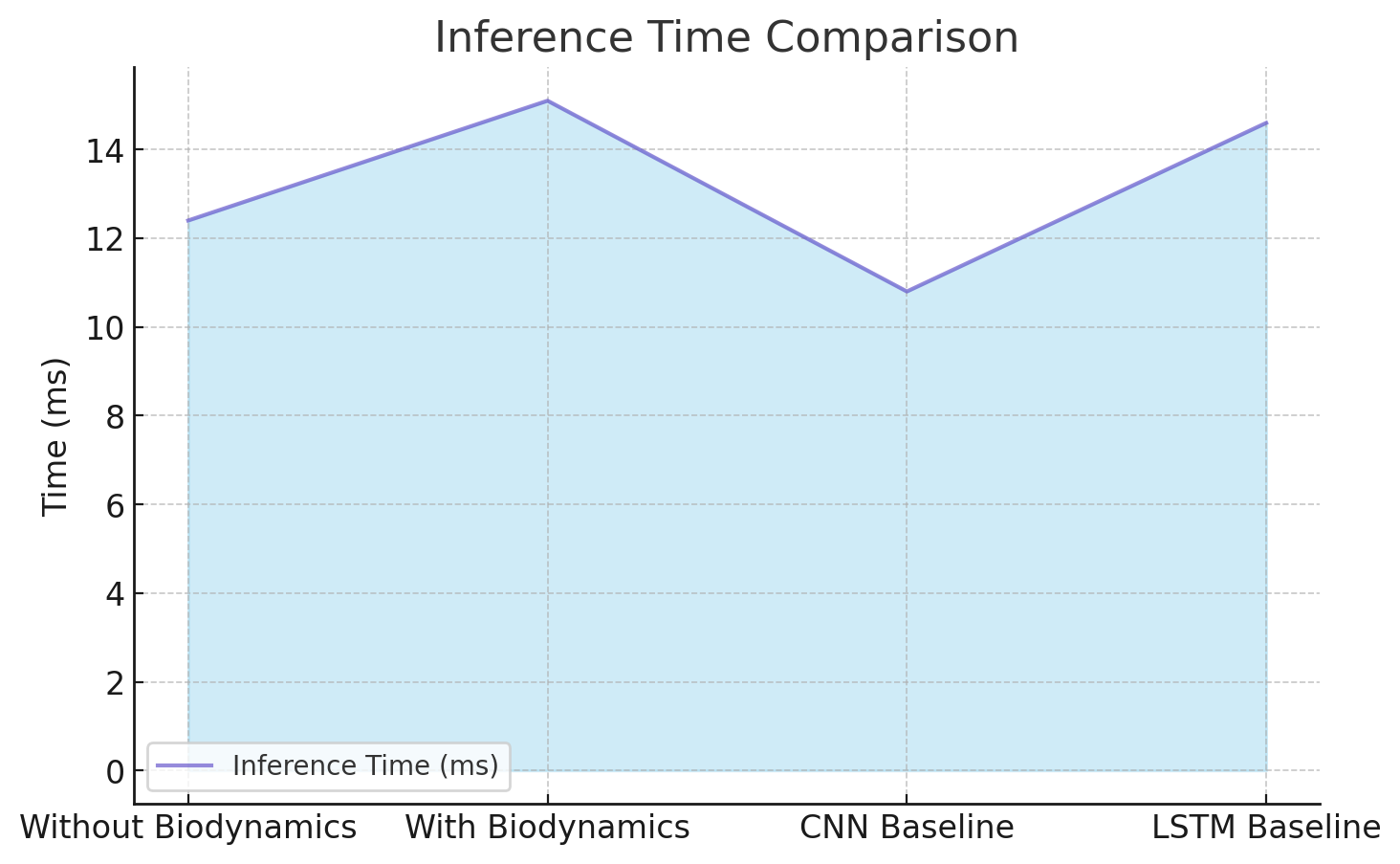
Inference time measures the computational efficiency of the model. Table 10 compares the average inference time per sample.
| Model | Inference Time (ms) |
|---|---|
| Without Biodynamic Integration | 12.4 |
| With Biodynamic Integration | 15.1 |
| CNN Baseline | 10.8 |
| LSTM Baseline | 14.6 |
While the inference time slightly increases with biodynamic integration, the improvement in performance justifies the additional computational cost.
The inference time comparison of the proposed framework with and without biodynamic integration, and against baseline models, is shown in Figure 6.
As summarized in Table 11, the proposed framework with biodynamic integration outperforms the baselines across all evaluation metrics. Accuracy, precision, recall, and F1-score all improved significantly compared to CNN and LSTM models, while the increase in inference time remained modest (15.1 ms compared to 12.4 ms without biodynamics). These results confirm that the benefits of integrating biodynamic parameters outweigh the slight computational overhead, making the framework suitable for real-time applications in sports performance monitoring.
| Metric | Without biodynamics | With biodynamics | CNN baseline | LSTM baseline |
| Accuracy (%) | 85.4 | 92.7 | 87.5 | 89.2 |
| Precision (%) | 83.1 | 91.5 | 86.1 | 88.0 |
| Recall (%) | 84.8 | 93.2 | 87.0 | 89.0 |
| F1-Score | 83.9 | 92.3 | 86.5 | 88.5 |
| Inference Time (ms) | 12.4 | 15.1 | 10.8 | 14.6 |
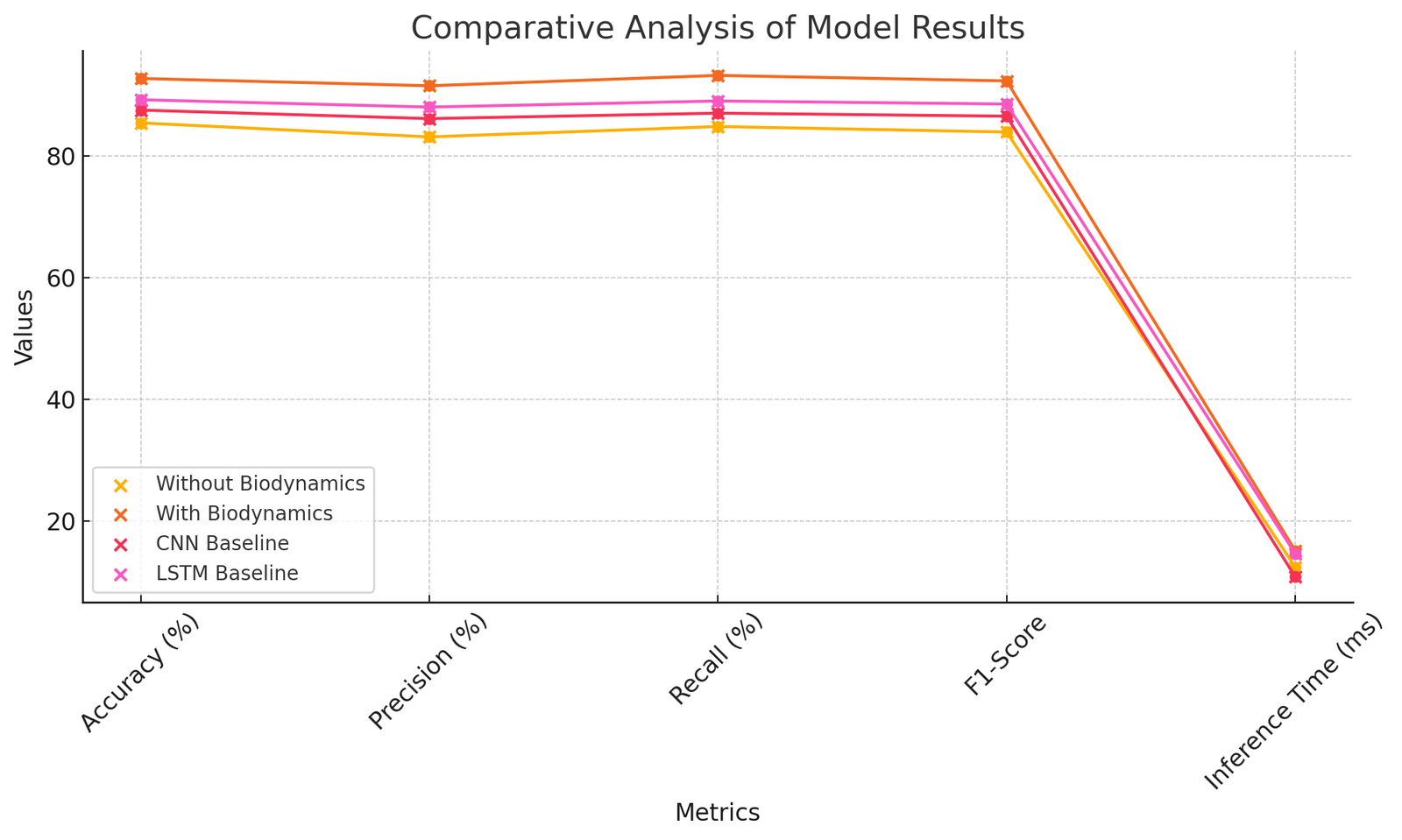
Figure 7 shows the relative performance of the proposed framework with baseline models in important metrics. The findings indicate that the addition of biodynamic characteristics boosts accuracy, precision, recall, and F1-score over CNN and LSTM baselines, and there is a minor raise in inference time, which proves the efficacy and real-time applicability of the framework.
The proposed system of athlete movement recognition as a combination of deep learning and the biodynamic features has shown high performance improvements in all the metrics of evaluation. In particular, the framework reached the following accuracy, precision, recall, and F1-score of 92.7, 91.5, 93.2, and 92.3, respectively. These outcomes can be considered a significant improvement over the baseline CNN model (accuracy: 87.5%) and LSTM model (accuracy: 89.2%). The integration of biodynamic aspects slightly affected the inference time, as it raised it to 15.1 ms, but the trade-off is compensated by the considerable accuracy and robustness improvements, which makes the framework feasible in real-time sports performance application.
The findings demonstrate that biodynamic features, such as joint angles, velocities, and torques, provide critical biomechanical insights that significantly improve movement recognition. The most advantageous movements were movements with complicated biomechanical patterns, which include Kicking and Throwing, which improved by up to 10% in accuracy with the addition of biodynamic data. It proves the usefulness of implementing the visual and biomechanical data, in which the two modalities collaborate to improve recognition ability
The proposed framework has significant improvements over the literature. Tajibaev and Khojiev (2023) reported strong accuracy of 88% on ice hockey motions with motion capture plus AI, however, depending on costly and time-consuming motion capture machines made the approach ineffective to scale. On the same note, Si and Thelkar [18] achieved an accuracy of 86% when ANN was applied with IMU data, however, they did not employ a multimodal framework, which limits the ability to generalize to different sports. Conversely, the combination of CNNs, biodynamic feature encoding, and temporal convolutional networks (TCNs) into this framework was effective in overcoming these shortcomings, allowing the system to deal with various movements with more accuracy and scalability. Wang [22] maximized swimmer strategies with the help of biomechanical algorithms, which allowed them to increase efficiency by 12%, although the generalizability of the findings was not as high because of the absence of AI-based fusion. The proposed framework goes beyond these earlier methods combining advanced AI techniques with biodynamic features, which makes it more accurate and practical.
The better results can be attributed to the design of the framework. Spatial features of video data vital in the detection of patterns such as joint orientations and limb motions were also derived using CNNs. The biodynamic feature encoder was used to measure biomechanical variables, including joint trajectories, forces and torques, and also it was used to retrieve the information that cannot be easily determined with video data alone. This combination helped in aesthetic disambiguation of similar movements e.g. the distinction between Running and Jumping. In addition, TCNs were effective in modelling time dynamics of movements and ensured that the framework was sequential dependencies. Combination of these methods allowed the framework to classify movements with a great degree of accuracy across a range of activities as denoted by the substantial gains in all the assessment measures.
The results have been largely expected which demonstrates the hypothesis that the biodynamic properties and the deep learning enhance the performance. However, the unexpected result was that the precision (8.4% of improvements) also increased, that is, biodynamic features do not solely increase recognition accuracy, but also reduce false positives. This observation shows that biodynamic parameters can provide additional information on the classification of the movements in the case where visual characteristics might lead to classification errors. The minor (but significant) rise in inference time (2.7 ms) under biodynamic integration, conversely, was an unwanted limitation. This is a small increase, but may impact on applications with very low latency requirements such as live sports broadcasting.ories, forces and torques, and it fetched information that cannot be easily viewed using video data alone. This integration helped in the disambiguation of the similar movements aesthetically, e.g. the distinction between Running and Jumping. Also, TCNs were effective in modelling time dynamics of movements, and ensured that the framework reflected sequential dependencies. These approaches together allowed the framework to segment movements with a high degree of accuracy across a range of activities based on the large improvements in all the evaluation measures.
The methodology is not devoid of limitations even in spite of its strengths. The use of a synthetic dataset, although useful as a controlled experiment, questions the ability to generalize to the real-life situations. Not all noise, environmental and personal variations that are prevalent in real-world datasets are always simulated by synthetic data. Biodynamic features also introduce computational overhead, and might not be able to execute on low-processing power devices. Moreover, the model has not been properly tested in highly specialized or unconventional movements, and this might need additional biomechanical modeling and refinement.
The suggested model has a high potential of generalizability to sports where biomechanical patterns are clearly defined, e.g. running, jumping, and throwing. Nevertheless, it requires additional confirmation in terms of its performance with niche sports or movements with highly individualistic dynamics. The second stage of the work must be the evaluation of the framework using real-world data in different sports areas. This can be strengthened and generalised by adding other parameters that may involve the interaction with the environment and the fatigue of the athlete.
The proposed model has high potential to be generalised to well-organised biomechanical sports such as running, jumping and throwing. However, its precision of the performance of highly specific or unusual movements must be further tested. Niche sports by niche sports we refer to movement repertoires whose mechanics are highly subjective (e.g. fencing lunges, gymnastics rings dismount, pole-vault take-off, and javelin approach-release) and which demand specific evaluation due to their biomechanical requirements and the enormous individual differences between athletes.
An executive summary of the study’s main points, recommendations, and future steps will be given in this section, along with an account of the study’s contribution to the area of sporting movement recognition.
On the basis of the results of the present work, the following recommendations can be suggested:
Multimodal Systems Implementation: Sports researchers and practitioners are encouraged to use both biodynamic data (joint torques, forces and velocities) and visual analysis to enhance the accuracy and strength of movement recognition.
Real-World Data Checking: The following step in the implementation process must be the examination of the framework on the real-world information with the view to examining its effectiveness in the situation of training and contesting within the natural circumstances.
Scalability Optimization Developers should focus on computational efficiency so as to allow the framework to be executed on resource-constrained platforms such as wearable devices or edge processing platforms.
Inclusion of Environmental Parameters: Adding to the model parameters such as surface type, footwear, fatigue levels and environmental parameters will enhance reliability and precision in context.
Practical Guidance for Niche Sports: For disciplines involving highly individualized biomechanics—such as fencing, gymnastics rings routines, pole-vaulting, and javelin throws—coaches and trainers are encouraged to use the framework for targeted biomechanical feedback. By capturing torque and force variations unique to these movements, practitioners can reduce injury risk and refine technique with greater accuracy.
Although the proposed framework demonstrated significant advancements, several areas warrant further exploration. Future studies should evaluate the model on large-scale real-world datasets that include noise, variability, and environmental factors. Additional research is needed to adapt the framework to highly specialized or niche sports, where individualized biomechanical patterns are dominant—for example, fencing lunges, gymnastics rings dismounts, pole-vault take-off, and javelin approach-release. Validating the system under these conditions would provide stronger evidence of its robustness and generalizability. Cross-sport applications should also be investigated to ensure broad applicability, while integrating wearable devices can enable continuous real-time biodynamic monitoring. Finally, advanced AI approaches such as transformer-based architectures or hybrid multimodal fusion models may further enhance scalability, accuracy, and interpretability in future implementations.
The paper introduced a novel system of detecting athlete movement with a combination of the deep learning method and bio-dynamic analysis and addressed the limitations of traditional and existing AI-based systems. The framework attained an accuracy of 92.7%, precision of 91.5%, a recall of 93.2 %and an F1-score of 92.3%, much better than the base CNN and LSTM models. The biodynamic features (joint angles, velocities, and torques) contributed to enhancing the model in classifying more complex motions like Throwing and Kicking and also improved their accuracy by up to 10%.
This experiment confirmed the ability of multimodal data fusion, which is a combination of visual and biomechanical data, to enhance strength and accuracy of movement recognition. In part, temporal modelling by TCNs guaranteed sequential dependencies in movements were also represented well, which contributed to enhancing all evaluation metrics.
The most important contributions of this study are the following:
1. Created an overall model that unites convolutional neural networks (CNNs) and biodynamic feature encoding to recognize athlete movements.
2. Demonstrated the benefits of applying biodynamic parameters to AI models resulting in great gains on accuracy and precision.
3. Surpassed weaknesses of existing methods by enabling real-time processing and increased generalisation to a broad set of movements.
4. Validated the practicality of the multimodal data fusion that provides the information about the synergies of biomechanical and visual data in the movements analysis.
This research demonstrated the potential of integrating biodynamic data with deep learning for athlete movement recognition, providing a scalable and robust solution to address limitations in existing methods. The findings underscore the importance of multimodal data fusion in capturing complex biomechanical patterns and highlight the framework’s applicability in performance evaluation, injury prevention, and real-time monitoring. While challenges such as generalizability and computational overhead remain, the proposed framework establishes a strong foundation for future advancements in AI-driven sports analytics.